A fantastic week at ESCMID Global 2026 in Munich! ✨🧬
At ESCMID Global 2026, many members of our group had the opportunity to share their ongoing projects with the gigantic clinical microbiology and infectious diseases community 🧫🌍. It was a great chance to present our work, exchange ideas with colleagues from different institutions, and discuss new approaches to understanding, detecting, and tackling challenges in applied microbiology. Also a nice time to have some fun in Munich!
Our contributions covered a broad range of topics: from antibiotic resistance mechanisms and bacterial interactions to rapid diagnostics, MALDI-TOF MS quality control, microbiome methods, diagnostic AI, and digital tools for susceptibility testing 💊🤖.
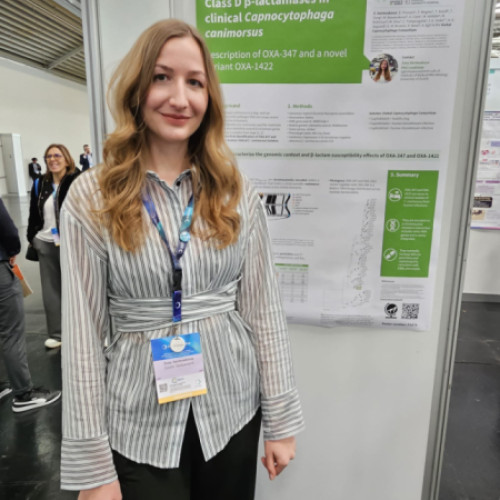
Zoey Germuskova — PhD candidate
Project: Antibiotic resistance in Capnocytophaga canimorsus
Zoey’s project investigates antibiotic resistance in Capnocytophaga canimorsus, a bacterium that can be transmitted from dogs and cats to humans and cause severe infections. The study identified β-lactamase enzymes OXA-347 and a novel variant, OXA-1422, in human clinical isolates, showing that they can reduce susceptibility to commonly used antibiotics such as penicillins and cephalosporins.
Mariella Greutmann — PhD candidate
Project: Bacterial interactions and antibiotic responses in multidrug-resistant E. coli
Mariella’s project uses pairwise co-culture experiments to study how bacterial interactions reshape antibiotic responses in multidrug-resistant E. coli. The results show that both bacterial growth and AMR gene expression vary depending on the antibiotic context and the neighbouring bacterial partner.
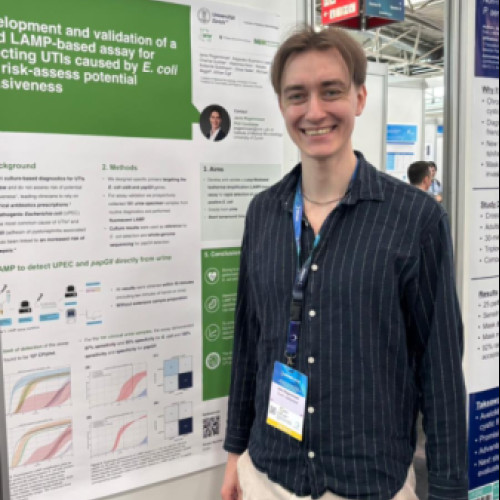
Janis Rogenmoser — PhD candidate
Project: Rapid LAMP-based detection of invasive-risk uropathogenic E. coli
Janis’s project developed and validated a rapid Loop-Mediated Isothermal Amplification (LAMP) assay to detect E. coli and the urosepsis-associated virulence marker papGII directly from urine samples. The assay delivered results within 40 minutes and showed strong diagnostic performance in routine clinical samples, highlighting its potential for early UPEC identification and invasiveness risk assessment.
Dr. Alejandro Guerrero-López — Postdoc
Project 1 : Knowledge distillation for MALDI-TOF antimicrobial resistance prediction
Alejandro’s knowledge distillation study shows that large AI models for MALDI-TOF antimicrobial resistance prediction can be compressed into much smaller neural networks without major loss of accuracy. Using a pretrained transformer as teacher and a lightweight student model, the project achieved near-teacher performance while substantially reducing model size, training time, and energy consumption, supporting more accessible and sustainable deployment of AI in microbiology.
Project 2: ASTAnnotator: digital measurement of disc diffusion inhibition zones
Alejandro also presented ASTAnnotator, an app supporting the digital measurement of antibiotic inhibition zones from disc diffusion images in a human-in-the-loop workflow. The application enables standardized, traceable, and offline annotation of routine AST plates, combining automatic disc detection with manual correction to improve reproducibility and facilitate retrospective analyses and future AI-assisted susceptibility testing workflows.
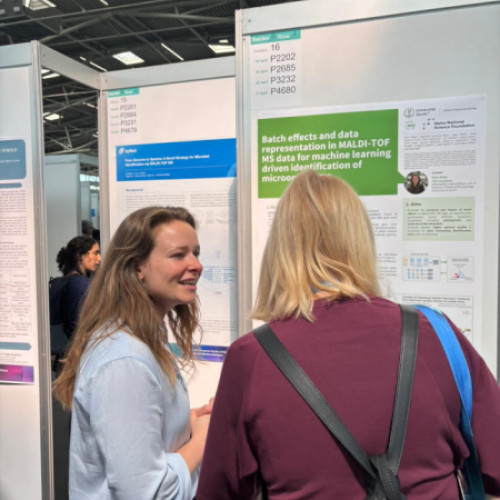
Eline Meijer — PhD candidate
Project: Batch effects and quality control in AI-driven MALDI-TOF MS analysis
Eline’s project investigates batch effects and quality control strategies in AI-driven MALDI-TOF MS analysis. The results demonstrate that machine-related batch effects can compromise identification beyond species level, while targeted quality control, particularly filtering spectra based on peak reproducibility, improves model performance across machines and supports more accurate downstream analyses.
Dr. Ashley Rooney — Postdoc
Project: DNA extraction methods for microbiome profiling in tissue biopsy samples
Ashley’s project compares DNA extraction methods for microbiome profiling in tissue biopsy samples. The study found that biopsy samples had low bacterial DNA yield, but that the extraction method did not significantly affect DNA yield, sequencing depth, pathogen detection, or microbiome richness.